Note
Click here to download the full example code
Tutorial for set prediction¶
In this tutorial, we propose set prediction for binary classification
estimated by MapieClassifier with the “lac”
method on two-dimensional dataset.
Throughout this tutorial, we will answer the following questions:
How does the number of classes in the prediction sets vary according to the significance level ?
Is the conformal method well calibrated ?
What are the pros and cons of the set prediction for binary classification in MAPIE ?
PLEASE NOTE: we don’t recommend using set prediction in MAPIE, even though
we offer this tutorial for those who might be interested.
Instead, we recommend the use of calibration (see more details in the
Calibration section of the documentation or by using the
CalibratedClassifierCV proposed by sklearn
or MapieCalibrator proposed in MAPIE).
from typing import List
import matplotlib.pyplot as plt
import numpy as np
from sklearn.calibration import CalibratedClassifierCV
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from mapie._typing import NDArray
from mapie.classification import MapieClassifier
from mapie.metrics import (classification_coverage_score,
classification_mean_width_score)
1. Conformal Prediction method using the softmax score of the true label¶
We will use MAPIE to estimate a prediction set such that
the probability that the true label of a new test point is included in the
prediction set is always higher than the target confidence level :
 .
We start by using the softmax score output by the base
classifier as the conformity score on a toy two-dimensional dataset.
We estimate the prediction sets as follows :
.
We start by using the softmax score output by the base
classifier as the conformity score on a toy two-dimensional dataset.
We estimate the prediction sets as follows :
First we generate a dataset with train, calibration and test, the model is fitted in the training set.
We set the conformal score
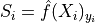 from the softmax output of the true class for each sample
in the calibration set.
from the softmax output of the true class for each sample
in the calibration set.Then we define
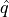 as being the
as being the
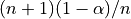 previous quantile of
previous quantile of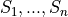  (this is essentially the
quantile
(this is essentially the
quantile 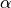 , but with a small sample correction).
, but with a small sample correction).Finally, for a new test data point (where
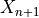 is known but
is known but
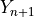 is not), create a prediction set
is not), create a prediction set
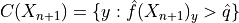 which includes
all the classes with a sufficiently high conformity score.
which includes
all the classes with a sufficiently high conformity score.
We use a two-dimensional dataset with two classes (i.e. YES or NO). The distribution of the data is a bivariate normal with arbitrary covariance matrices for each label.
centers = [(-2, 0), (2, 0)]
covs = [np.array([[2, 1], [1, 2]]), np.diag([4, 1])]
x_min, x_max, y_min, y_max, step = -6, 8, -6, 8, 0.1
n_samples = 2000
n_classes = 2
np.random.seed(42)
X = np.vstack(
[
np.random.multivariate_normal(center, cov, n_samples)
for center, cov in zip(centers, covs)
]
)
y = np.hstack([np.full(n_samples, i) for i in range(n_classes)])
X, X_val, y, y_val = train_test_split(X, y, test_size=0.5)
X_train, X_cal, y_train, y_cal = train_test_split(X, y, test_size=0.3)
X_c1, X_c2, y_c1, y_c2 = train_test_split(X_cal, y_cal, test_size=0.5)
xx, yy = np.meshgrid(
np.arange(x_min, x_max, step), np.arange(x_min, x_max, step)
)
X_test = np.stack([xx.ravel(), yy.ravel()], axis=1)
Let’s see our training data
colors = {0: "#1f77b4", 1: "#ff7f0e"}
y_train_col = list(map(colors.get, y_train))
fig = plt.figure()
plt.scatter(
X_train[:, 0],
X_train[:, 1],
color=y_train_col,
marker="o",
s=10,
edgecolor="k",
)
plt.xlabel("X")
plt.ylabel("Y")
plt.show()
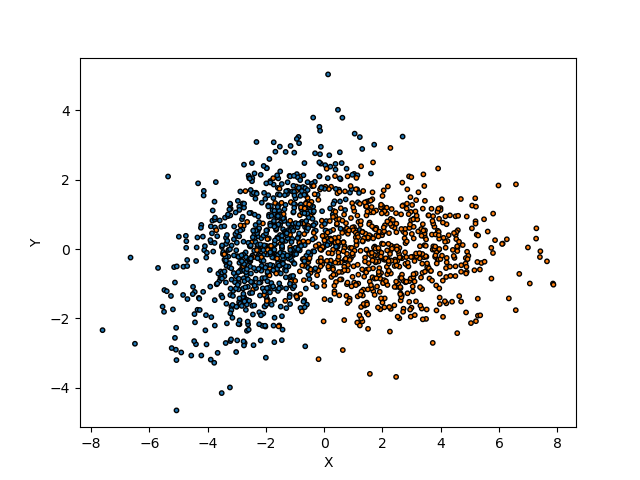
We fit our training data with a Gaussian Naive Base estimator.
We first apply a probability calibration with
CalibratedClassifierCV proposed by sklearn
so that scores can be interpreted as probabilities
(see documentation for more information).
Then we apply MapieClassifier in the
calibration data with the methods score
to the estimator indicating that it has already been fitted with
cv=”prefit”.
We then estimate the prediction sets with differents alpha values with a
fit and predict process.
clf = GaussianNB().fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_pred_proba = clf.predict_proba(X_test)
y_pred_proba_max = np.max(y_pred_proba, axis=1)
calib = CalibratedClassifierCV(
estimator=clf, method='sigmoid', cv='prefit'
)
calib.fit(X_c1, y_c1)
mapie_clf = MapieClassifier(
estimator=calib, method='lac', cv='prefit', random_state=42
)
mapie_clf.fit(X_c2, y_c2)
alpha = [0.2, 0.1, 0.05]
y_pred_mapie, y_ps_mapie = mapie_clf.predict(
X_test, alpha=alpha,
)
MAPIE produces two outputs:
y_pred_mapie: the prediction in the test set given by the base estimator.y_ps_mapie: the prediction sets estimated by MAPIE using the “lac” method.
Let’s now visualize the distribution of the conformity scores with the two methods with the calculated quantiles for the three alpha values.
def plot_scores(
alphas: List[float],
scores: NDArray,
quantiles: NDArray,
method: str,
ax: plt.Axes,
) -> None:
colors = {0: "#1f77b4", 1: "#ff7f0e", 2: "#2ca02c"}
ax.hist(scores, bins="auto")
i = 0
for quantile in quantiles:
ax.vlines(
x=quantile,
ymin=0,
ymax=100,
color=colors[i],
linestyles="dashed",
label=f"alpha = {alphas[i]}",
)
i = i + 1
ax.set_title(f"Distribution of scores for '{method}' method")
ax.legend()
ax.set_xlabel("scores")
ax.set_ylabel("count")
fig, axs = plt.subplots(1, 1, figsize=(10, 5))
conformity_scores = mapie_clf.conformity_scores_
quantiles = mapie_clf.quantiles_
plot_scores(alpha, conformity_scores, quantiles, 'lac', axs)
plt.show()
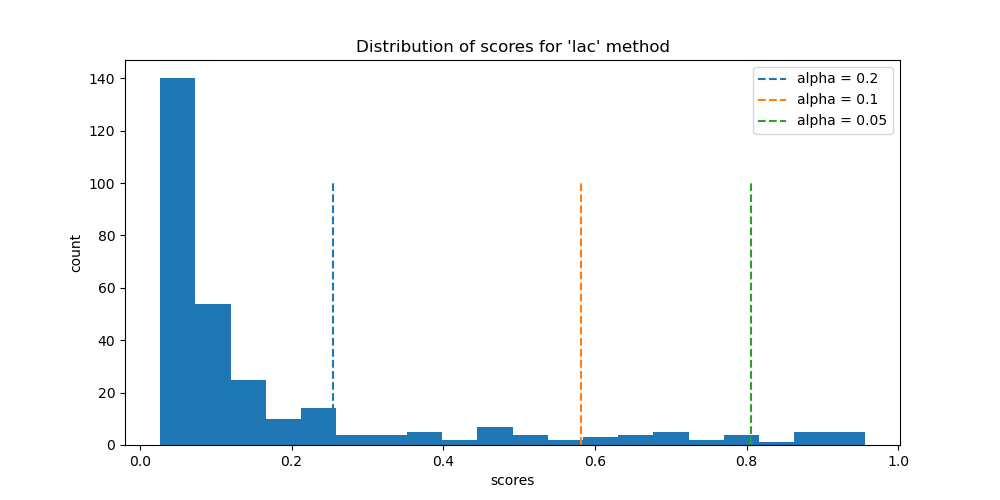
We will now compare the differences between the prediction sets of the different values of alpha.
def plot_prediction_decision(y_pred_mapie: NDArray, ax) -> None:
y_pred_col = list(map(colors.get, y_pred_mapie))
ax.scatter(
X_test[:, 0],
X_test[:, 1],
color=y_pred_col,
marker=".",
s=10,
alpha=0.4,
)
ax.scatter(
X_train[:, 0],
X_train[:, 1],
color=y_train_col,
marker="o",
s=10,
edgecolor="k",
)
ax.set_title("Predicted labels")
def plot_prediction_set(y_ps: NDArray, alpha_: float, ax) -> None:
tab10 = plt.cm.get_cmap("Purples", 4)
y_pi_sums = y_ps.sum(axis=1)
num_labels = ax.scatter(
X_test[:, 0],
X_test[:, 1],
c=y_pi_sums,
marker="o",
s=10,
alpha=1,
cmap=tab10,
vmin=0,
vmax=3,
)
ax.scatter(
X_train[:, 0],
X_train[:, 1],
color=y_train_col,
marker="o",
s=10,
edgecolor="k",
)
ax.set_title(f"Number of labels for alpha={alpha_}")
plt.colorbar(num_labels, ax=ax)
def plot_results(
alphas: List[float], y_pred_mapie: NDArray, y_ps_mapie: NDArray
) -> None:
_, [[ax1, ax2], [ax3, ax4]] = plt.subplots(2, 2, figsize=(10, 10))
axs = {0: ax1, 1: ax2, 2: ax3, 3: ax4}
plot_prediction_decision(y_pred_mapie, axs[0])
for i, alpha_ in enumerate(alphas):
plot_prediction_set(y_ps_mapie[:, :, i], alpha_, axs[i+1])
plt.show()
plot_results(alpha, y_pred_mapie, y_ps_mapie)
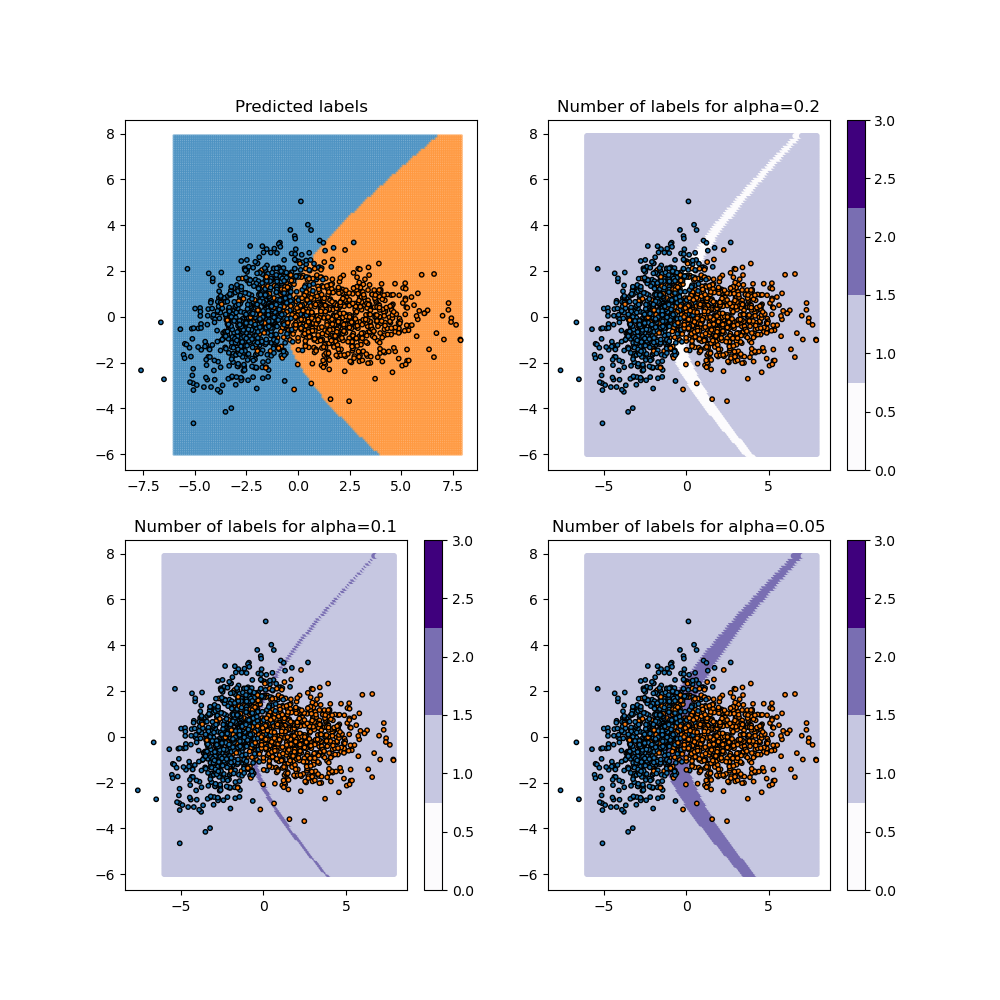
For the “lac” method, when the class coverage is not large enough, the prediction sets can be empty when the model is uncertain at the border between two classes. These null regions disappear for larger class coverages but ambiguous classification regions arise with both classes included in the prediction sets.
In other words, the choice of a class coverage leads to an associated prediction decision and vice versa.
A remarkable case: if our prediction decision is based on a threshold of 0.5, all prediction sets contain only one class (because binary classification). There are no ambiguous or uncertain classification regions. We’ll illustrate this later. Therefore, the accuracy of the model is similar to its coverage.
print(
f"Accuracy of the model with 'lac' method: "
f"{100*np.mean(mapie_clf.predict(X_val) == y_val)}%"
)
Out:
Accuracy of the model with 'lac' method: 89.1%
Let’s now compare the effective coverage and the average of prediction set
widths as function of the 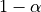 target coverage.
target coverage.
alpha_ = np.arange(0.02, 0.98, 0.02)
calib = CalibratedClassifierCV(
estimator=clf, method='sigmoid', cv='prefit'
)
calib.fit(X_c1, y_c1)
mapie_clf = MapieClassifier(
estimator=calib, method='lac', cv='prefit', random_state=42
)
mapie_clf.fit(X_c2, y_c2)
_, y_ps_mapie = mapie_clf.predict(
X, alpha=alpha_
)
coverage = np.array([
classification_coverage_score(y, y_ps_mapie[:, :, i])
for i, _ in enumerate(alpha_)
])
mean_width = [
classification_mean_width_score(y_ps_mapie[:, :, i])
for i, _ in enumerate(alpha_)
]
def plot_coverages_widths(alpha, coverage, width, method):
_, axs = plt.subplots(1, 3, figsize=(15, 5))
axs[0].set_xlabel("1 - alpha")
axs[0].set_ylabel("Quantile")
axs[0].scatter(1 - alpha, mapie_clf.quantiles_, label=method)
axs[0].legend()
axs[1].scatter(1 - alpha, coverage, label=method)
axs[1].set_xlabel("1 - alpha")
axs[1].set_ylabel("Coverage score")
axs[1].plot([0, 1], [0, 1], label="x=y", color="black")
axs[1].legend()
axs[2].scatter(1 - alpha, width, label=method)
axs[2].set_xlabel("1 - alpha")
axs[2].set_ylabel("Average size of prediction sets")
axs[2].legend()
plt.show()
plot_coverages_widths(alpha_, coverage, mean_width, 'lac')
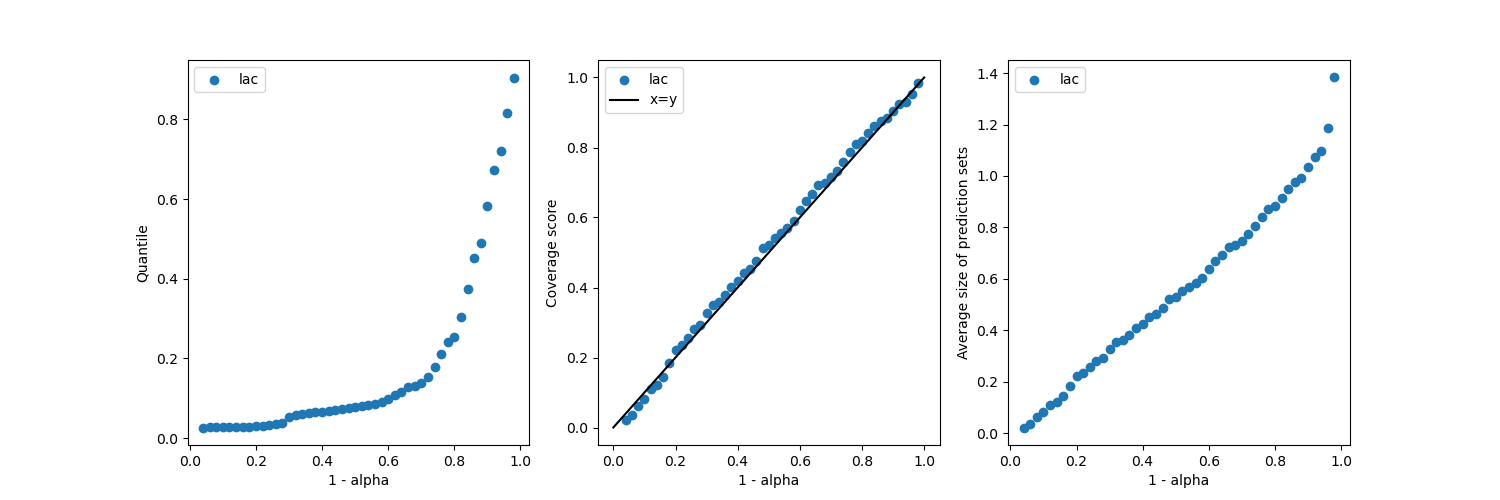
It is seen that the method gives coverages close to the target coverages,
regardless of the 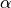 value.
value.
alpha_ = np.arange(0.02, 0.16, 0.01)
calib = CalibratedClassifierCV(
estimator=clf, method='sigmoid', cv='prefit'
)
calib.fit(X_c1, y_c1)
mapie_clf = MapieClassifier(
estimator=calib, method='lac', cv='prefit', random_state=42
)
mapie_clf.fit(X_c2, y_c2)
_, y_ps_mapie = mapie_clf.predict(
X, alpha=alpha_
)
non_empty = np.mean(
np.any(mapie_clf.predict(X_test, alpha=alpha_)[1], axis=1), axis=0
)
idx = np.argwhere(non_empty < 1)[0, 0]
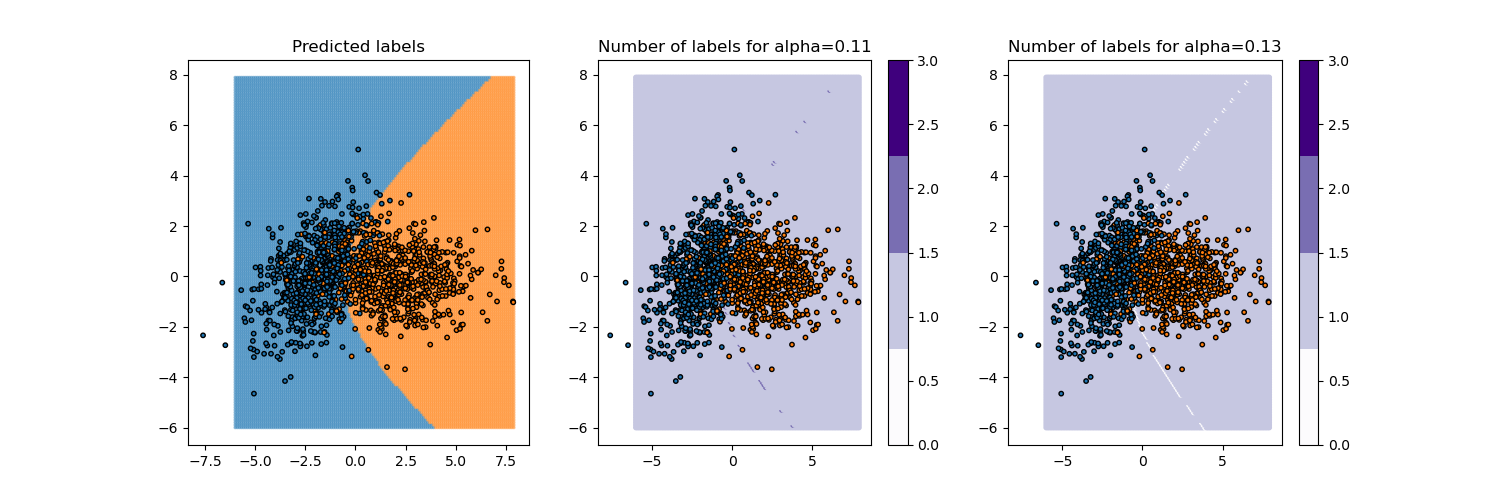
_, axs = plt.subplots(1, 3, figsize=(15, 5))
plot_prediction_decision(y_pred_mapie, axs[0])
_, y_ps = mapie_clf.predict(X_test, alpha=alpha_[idx-1])
plot_prediction_set(y_ps[:, :, 0], np.round(alpha_[idx-1], 3), axs[1])
_, y_ps = mapie_clf.predict(X_test, alpha=alpha_[idx+1])
plot_prediction_set(y_ps[:, :, 0], np.round(alpha_[idx+1], 3), axs[2])
plt.show()
Total running time of the script: ( 0 minutes 3.206 seconds)